iRODS is a decentralized virtual file system that is separate from your normal directories in Linux or Windows. In iRODS, directories are called collections and they can contain further sub-collections (sub-directories) or data objects (files). In practice, you can treat collections and data as directories and files for the sake of simplicity. Files and folders in your own local and mounted directories can be archived (copied) to the remote iRODS file system provided by the ZDV. Archiving is done by first creating a directory (collection) in the remote path and then copying your local files and directories to the remote path in iRODS. The archived files can be copied back or downloaded years later. The archives can be private and only stored long-term, they can be restricted to research groups or offered for public download.
Archiving in iRODS is currently possible via the command line of the ZDV Linux work PCs, the web server linux.zdv.uni-mainz.de and the supercomputer MOGON II/MOGON-NHR. MOGON also has a special iRODS wiki. In the future, there will be new and more user-friendly options for accessing iRODS. Currently, only users with a university account can use iRODS. If your current work computer does not have a standard ZDV Linux installation, you can access the Linux servers via SSH. From there, you can use the available iRODS iCommands to archive data. Personal computers with an iRODS iCommand client installation can also access the ZDV iRODS archive.
Before accessing the iRODS server, a configuration file must be available and you must authenticate yourself with your JGU account. The configuration file “irods_environment.json” is located in the hidden folder “.irods” and has the following content:
{
"irods_authentication_scheme": "pam_password",
"irods_client_server_negotiation": "request_server_negotiation",
"irods_client_server_policy": "CS_NEG_REQUIRE",
"irods_encryption_algorithm": "AES-256-CBC",
"irods_encryption_key_size": 32,
"irods_encryption_num_hash_rounds": 16,
"irods_encryption_salt_size": 8,
"irods_host": "irods.zdv.uni-mainz.de",
"irods_port": 1247,
"irods_user_name": "FILL IN YOUR USERNAME HERE",
"irods_home": "/zdv/home/FILL IN YOUR USERNAME HERE",
"irods_zone_name": "zdv"
}Do not forget to enter your correct user name. Then enter the command “iinit” and your JGU password.
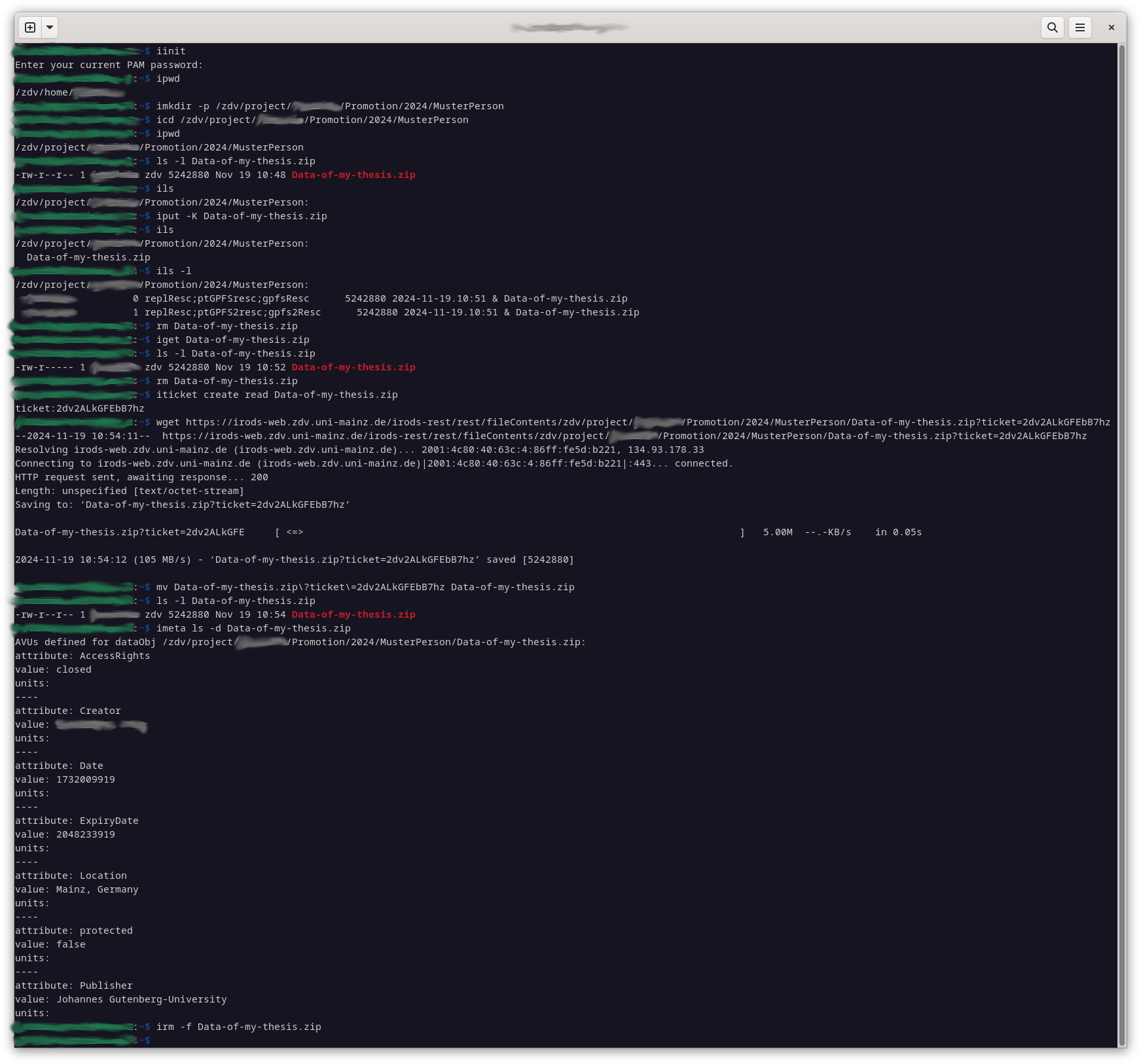
Here is a short summary of the most important iRODS commands with some important command line parameters. Note that they start with an ‘ i ‘:
| Command | Parameters | Description |
|---|---|---|
ipwd | Print current iRODS working directory (collection) | |
ils | -l, -L, -A, -r | Lists of the iRODS directory (collection) (-l: with details; -L: more details; -A: ACL, -r recursive) |
icd | Change iRODS directory (collection) to the target path | |
imkdir | -p | create a new directory (collection) (directory; -p: create full path with parents) |
iput | -K, -r, –metadata | Upload files/directories, (K: calculate and validate checksums; -r: recursive, –metadata add descriptions of the data: “Title;My data;;Description;Research data;;”) |
iget | -r, -f | Download iRODS target data to the current local path, -r recursive, -f overwrite |
iticket | create read | Create a ticket for downloading files from the Internet |
imeta | ls -d/-C | Listing the metadata of an iRODS file or directory (collection) |
imeta | set -d/-C Key “Value” | Add/update metadata to an iRODS file or directory (collection); e.g: Key = Title, “Value” = My research data: Key = Title, “Value” = My research data |
Here is a brief summary of our iRODS wrapper commands that simplify the archiving process of research data. Note that wrappers start with an ‘ i_ ‘, unlike pure iRODS commands which start with ‘i’:
| Command | Parameters | Description |
|---|---|---|
source /usr/local/bin/i_init.sh | For linux.zdv.uni-mainz.de | Initializes iRODS and displays the current remote iRODS directory after each subsequent command. |
i_exit | Deletes the current iRODS remote directory and cleans up the local settings. | |
i_archive | Uploads the 1st argument to the current remote iRODS directory/collection and adds the metadata of the .json file of the second argument to all uploaded files. | |
i_metaupdate | Updates the (first argument) remote iRODS directory/collection and updates the metadata (second argument) using the .json file. | |
i_publish | Creates an iRODS web download ticket and links for all files in the path (first argument). Anyone can use the links to download the files. | |
i_ticketget | Prints the iRODS ticket of a remote iRODS path, if available. | |
i_downlinkget | Uses the iRODS ticket and prints the public web download links. |
A simplified procedure for archiving research data would use the simplified wrappers and look as follows:
source /usr/local/bin/i_init.sh for the systems linux.zdv.uni-mainz.de
Provides the wrapper commands and initializes iRODS
imkdir -p /zdv/home/MUSTERPERSON/project123/
creates a new remote iRODS path to which the data is copied;
icd /zdv/home/MUSTERPERSON/project123/
changes the current remote iRODS directory to the newly created one;
i_archive /fullpath/some.file metadata.json
archives a local file some.file into the current remote iRODS path by copying it and transferring the metadata from the metadata.json files.
i_publish /zdv/home/MUSTERPERSON/project123/some.file
creates a ticket number (e.g.: ABCDEFG123456890) and prints the generated web download link.
Metadata is accepted by iRODS as triplets (Attribute [i.e: Title], Value [i.e: “Research Data from X publication”],, Unit[meist immer leer gelassen]).
The first two fields Attribute and Value are mandatory fields and must not be empty, the Unit is optional.
You can manually add metadata to each remote iRODS file and directory after uploading by selectingimeta set -d some.file Title "My research data from Publication X"
This would be very complex and time-consuming, so it is advisable to define a .json metadata file for all files and directories of a single archive tree.
i_archive some.file metadata.json
Can be used to upload your local files with metadata all at once.
i_metaupdate /zdv/home/MUSTERPERSON/project123/some.file metadata.json
Can be used to update a single remote file/directory with metadata.
This flat metadata.json example requires a minimal set of metadata attributes:
{
"Title":"",
"ResourceType":"",
"Project":"",
"Keywords":"",
}
{
"Title":"My Scientific Data from XYZ publication",
"ResourceType":"Tables, Texts, Images",
"Project":"BMBF-12345, DFG-67890",
"Keywords":"Thermodynamics, Simulation, HPC, MPI , XYZ, ,BMBF, DFG",
}
The following attributes are set automatically when uploading files/directories and are not required in the .json file:
Creator, Publisher, Location, Date, ExpiryDate(Date + 10 years), protected (default: "false") .
The following attributes are recommended in your .json metadata file:
{
"Title":"",
"ResourceType":"",
"Project":"",
"Keywords":"",
"Contributor":"",
"Reference":"",
"License":""
}
{
"Title":"My Scientific Data from XYZ publication",
"ResourceType":"Tables, Texts, Images",
"Project":"BMBF-12345, DFG-67890",
"Keywords":"Thermodynamics, Simulation, HPC, MPI , XYZ, ,BMBF, DFG",
"Contributor":"Co-author1, Co-author2, Co-author3",
"Reference":"",
"License":"GPLX, CC0, CC-BY"
}
You can confirm that the metadata has been set by using:imeta ls -d /zdv/home/MUSTERPERSON/project123/some.file for filesimeta ls -C /zdv/home/MUSTERPERSON/project123/ for directories.
Summary of imeta:
| Parameters | Description |
|---|---|
| add|set|rm|ls|cp | command, see the next table for details(ls|cp do not need the AVU triplet) |
| -d dataObject |-C directory/collection | which object/collection (file/path) is to be queried/edited |
| Attributes Value [Unit] | AVU Triplet, in which the unit is optional |
Command description:
| Command | Description |
|---|---|
| add | add an AV(U) triplet |
| set | Set a single value |
| rm | Remove an AV(U) triplet |
| ls | List existing metadata. If attribute is specified, only metadata of the specified attribute is listed |
| cp | Copy existing metadata.
Requires a destination and a source (e.g. imeta cp -d source -c target) |
Publishing research data:
A ticket for downloading files must be created for public access.
With this ticket and the path to the remote iRODS storage location, anyone can download the information and content.
Already archived data is published with :
i_publish
i_publish /zdv/home/MUSTERPERSON/project123/some.file
generates a ticket number (i.e: ABCDEFG123456890) and prints the generated web download link
You can also perform the publishing process manually using iRODS commands instead:iticket create read /zdv/home/MUSTERPERSON/project123/some.file
generates a ticket number (i.e: ABCDEFG123456890) udie is used to download some.file from the network
Then you need to add the path so that it looks like this: wget https://irods-web.zdv.uni-mainz.de/irods-rest/rest/fileContents/zdv/home/MUSTERPERSON/some.file?ticket=ABCDEFG123456890Data Policy and Licensing
The “creator” is responsible for ensuring that the further use of third-party data is lawful (copyright law) and that personal data is handled correctly (GDPR).
This applies to all archived data, even if the “creator” is no longer employed at the university or if the data is not public.
This decision aid can help you to decide whether your data may be published.
If your data can actually be published, there are different types of licenses for different cases:
- Software
- Art, pictures, text, etc.
- Data records
The applicability of CC-BY licenses for software is not recommended: see CC recommendation and discussion. The same applies to datasets where publication under a CC licence other than CC0 is questionable. Other licenses for datasets can be found at Open Definition Licenses Service.
Avoid proprietary file formats, as you don’t know if the software to open them will still be available in a few years’ time. Try to use open standards.
You can contact unix@uni-mainz.de to make an appointment with our technical support team.
They will be able to help clarify any further questions you may have and guide you through the process.
Archiving is intended for research data that is part of a previously published paper. Archived data is intended to be used to replicate research results. It may also be published for use by others, or kept private to meet funding requirements. If you would like to backup a lab PC, please go to: https://www.en-zdv.uni-mainz.de/data-backup/
To archive research data, you don’t need to change the way you do research or any of your existing data workflows. You can archive data at any time during the research life cycle. Nevertheless, it it recommended to plan ahead, which data should be archived with which metadata. For this we also provide the Research Data Management Organizer (RDMO).
Data can be archived at any time by creating relevant metadata and technical metadata and then uploading it to the archive. Metadata summarises key information about the research, such as title, description, type, format, licence, keywords, contributors, references, etc. Technical metadata describes what the files contain, how they are used in research, how they relate to each other and how they can be edited. All of this information is useful for others to find and understand the research data.
All research data of a single publication to be archived should be collected in a single parent directory. The data within the parent directory can then be sorted into different directories based on a logical hierarchy chosen by the authors. The metadata of the collected data should then be defined, i.e. title, description, type, format, license, keywords, contributors, reference, etc. Technical descriptions of the research data (technical metadata) should also be prepared and included in the parent directory as human-readable files such as “.txt”. This technical metadata should describe what the contents of the files represent, how they are used in research, how they relate to each other, and how they can be processed. The parent directory should be compressed into an open format if storage requirements allow.
Once you have collected your research data and tagged it with metadata, you can follow these steps [link] to archive it.
Several large files can be archived individually, while large quantities of small files belonging to the same data set should be compressed in a single archive file (zip, tar, rar). If possible, all files of the same data set should be combined in a single archived (compressed) file. This makes it easier to find complete data sets that are to be distributed together.
The data should be available to be copied from either:
- a Linux computer installation provided by the ZDV [link]
- the Mogon HPC system at JGU [link]
- any computer with connected JGU group storage network drives [link]
- a personal computer with Kerberos client authentication packages installed
- any computer with SSH access to linux.zdv.uni-mainz.de [link] or the MOGON HPC system [link]
- a Windows PC that uses the Linux subsystems or Linux virtual machines.
The source code needed to process, generate or understand your research data should be provided with the data itself whenever it is available or possible. The archive itself is intended for long-term, immutable data
By default, the data is archived for 10 years after uploading.
For archiving datasets over 1TB, it is recommended to contact us at zdv-forschungsdaten@uni-mainz.de to help you ensure smooth performance and stability during the archiving process.
There is currently no storage limit for the amount of datasets to be archived, but individual files or compressed datasets should not be larger than ~8TB.
If the archived data sets are to be retrieved via HTTP interfaces, a maximum size of 5 GB per file is recommended.
The capacity limitations are only given by the storage location of the original data to be copied, i.e. 100 GB at home on the MOGON systems.
There is currently no limit to the number of files to be archived. However, if the data will be tranfered to the tape library it might be impossible to retrieve millilios of small files. Therefore, if possible, all files of the same data set should be combined in a compressed file. This makes it easier to find a complete data set that should belong together.
To give other users access to your research data archive, you must carry out the following steps [Link].
Your research data can be published during the archiving process or afterwards in the following steps [Link].
First, metadata must be captured, such as title, description, type, format, license, keywords, contributor, reference, etc. Once you have defined these and other helpful descriptive fields, you can follow these steps [Link] to add metadata.
Technical metadata must be created by the owner of the research data. Technical descriptions of the research data should then be included as human-readable files such as “.txt” alongside the normal data. This technical metadata should describe what the content of the files represents, how they are used in the research field, how they relate to each other and how they can be processed.
Under normal circumstances, the archive is intended to retain unalterable data for decades. However, the archived research data may be updated in certain cases to add or change existing files. This may be the case if incorrect data is inadvertently uploaded or if the underlying published research changes. Research data that has been archived long-term to meet government or financial requirements cannot and should not be deleted.
iRODS uses “resources” to archive the collections (directories) and data objects (files). The resources are organized hierarchically. The iRODS archive of the ZDV currently has a replication ressource consisting of two GPFS-Filesystem at diffenrent locations. When you archive data, success is only reported, if both copies are created. In the future one copy of the data might be migrated to the tape library.